Genetic Circuit Could Help Engineered Bacteria Hunt Down Tumors
New research from Columbia Engineering could help make sure microbial therapies target disease sites, not the rest of the body
ABOUT THE STUDY
The study is titled "Enhancing the tropism of bacteria via genetically programmed biosensors."
The study appeared in the journal Nature Biomedical Engineering on July 29, 2021.
Authors are: Tiffany Chien, Tetsuhiro Harimoto, Benjamin Kepecs , Kelsey Gray, Courtney Coker, Nicholas Hou, Kelly Pu, Tamjeed Azad, Andoni Nolasco, Martina Pavlicova and Tal Danino.
Department of Biomedical Engineering, Columbia University
Biostatistics Department, Columbia University
Data Science Institute, Columbia University
Herbert Irving Comprehensive Cancer Center, Columbia University
This work was supported by the DoD Idea Development Award (LC160314), DoD Era of Hope Scholar Award (BC160541), Honjo International Foundation Scholarship and NIH F99CA253756.
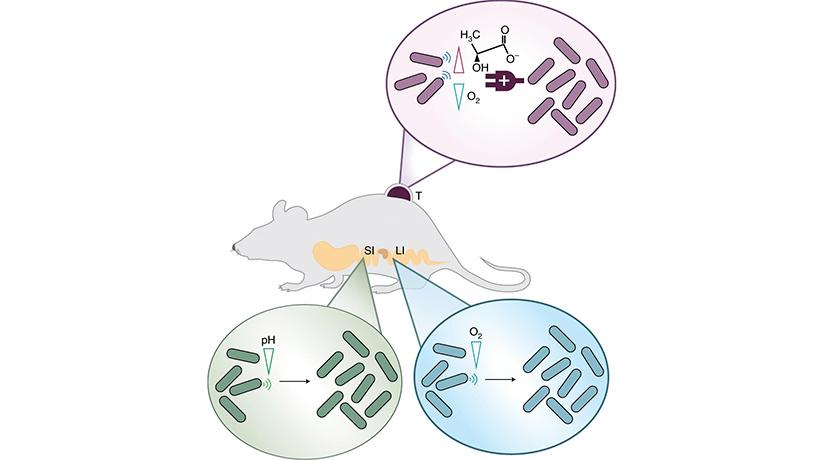
Schematic of biosensors that genetically engineered bacteria can use to stay at diseased sites. (Top) To stay at tumors, the microbes are engineered to favor high lactate and low oxygen levels. (Bottom left) The small intestine favors low pH conditions. (Bottom right) The large intestine favors low oxygen conditions.
New York, NY—August 2, 2021— Genetically engineered bacteria might one day find use in therapeutic applications, growing within the human body to sense and respond to problems such as inflammation, infections, or cancer. Now scientists at Columbia Engineering are designing ways to help confine these cells at diseased tissues or regions where they can do the most good, and not travel elsewhere where they may do harm.
A new study detailing this work, titled "Enhancing the tropism of bacteria via genetically programmed biosensors," appeared July 29 in the journal Nature Biomedical Engineering. The study was co-led by Tiffany Chien and Tetsuhiro Harimoto, PhD candidates in the lab of Tal Danino, associate professor of biomedical engineering at Columbia Engineering.
A major challenge in using engineered microbes for medicine is safety, "because they may grow in unintended body sites, causing toxicity," Chien said. To address this problem, the researchers genetically engineered bacteria that can sense cues in their environment connected with cancer and selectively grow only in those sites. One such hallmark was low oxygen, as the rampant cell growth, abnormal blood vessels and altered metabolism seen with tumors often reduces oxygen levels. Another was the molecule lactate, which cancer cells often produce in high amounts. If the bacteria strayed to areas without these cues, essential genes of theirs shut down, preventing them from growing in these sites.
However, some parts of the body may share certain traits with tumors—for instance, the large intestine also has low oxygen levels. To help prevent therapeutic bacteria from growing in healthy areas that are similar to diseased ones in some respects, the scientists created a "genetic circuit" so the bacteria would only grow in sites with, say, both low oxygen and high lactate levels. "We demonstrated that this approach improved tumor specificity of the bacteria when administered to tumor-bearing mice," Harimoto said.
The study also showed containment of the probiotic bacteria in the gastrointestinal tract of mice using engineered pH or oxygen dependent biosensors. "It is great to see collaboration between students in the biomedical engineering department, addressing an important problem towards clinical translation of bacterial cancer therapy," Danino said.
###
Columbia Engineering
Columbia Engineering, based in New York City, is one of the top engineering schools in the U.S. and one of the oldest in the nation. Also known as The Fu Foundation School of Engineering and Applied Science, the School expands knowledge and advances technology through the pioneering research of its more than 220 faculty, while educating undergraduate and graduate students in a collaborative environment to become leaders informed by a firm foundation in engineering. The School's faculty are at the center of the University's cross-disciplinary research, contributing to the Data Science Institute, Earth Institute, Zuckerman Mind Brain Behavior Institute, Precision Medicine Initiative, and the Columbia Nano Initiative. Guided by its strategic vision, "Columbia Engineering for Humanity," the School aims to translate ideas into innovations that foster a sustainable, healthy, secure, connected, and creative humanity.